We study how biological architecture can be read, engineered, and functionally translated across scales. Our research integrates molecular imaging pipelines for high-dimensional spatial biology with the engineering of neuronal architectures into functional bioelectronic and neurodevice platforms.
Integrated Molecular Imaging Pipeline
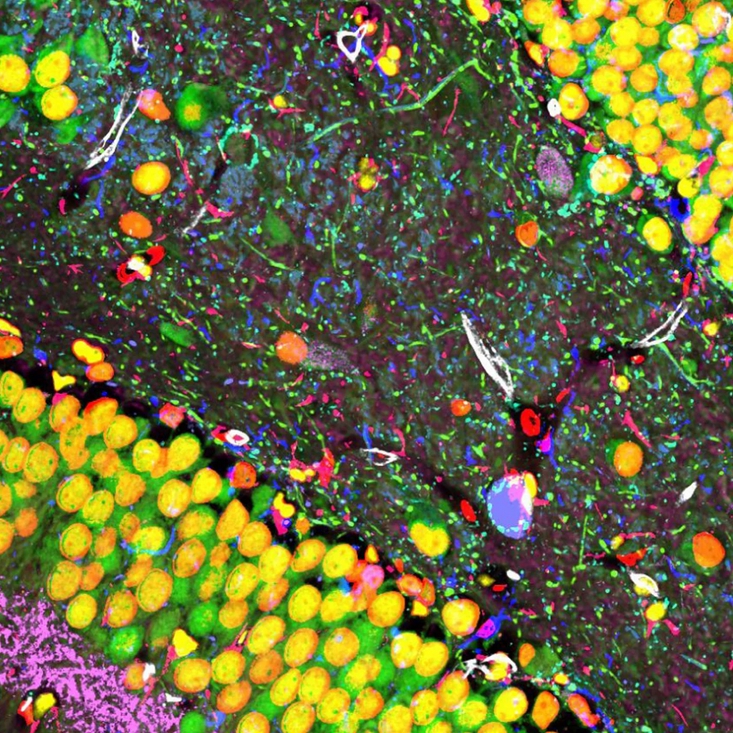
We develop integrated molecular imaging pipelines for the quantitative analysis of cells and tissues. Rather than focusing on imaging alone, we optimize the full experimental and analytical workflow, including tissue preparation, labeling chemistry, iterative processing, microscopy, image alignment, signal extraction, and downstream quantitative analysis. This systems-level approach improves reproducibility, scalability, and comparability across samples and experiments.
Our goal is to make molecular imaging more accurate and more useful as a quantitative platform for biology. Using multiplexed immunofluorescence, spectral unmixing-free imaging, cyclic imaging, and computational image registration, we generate high-dimensional spatial datasets that reveal molecular states, cellular organization, and tissue architecture in cancer, the brain, and other complex biological systems.
Imaging Chemistry and Expansion Engineering
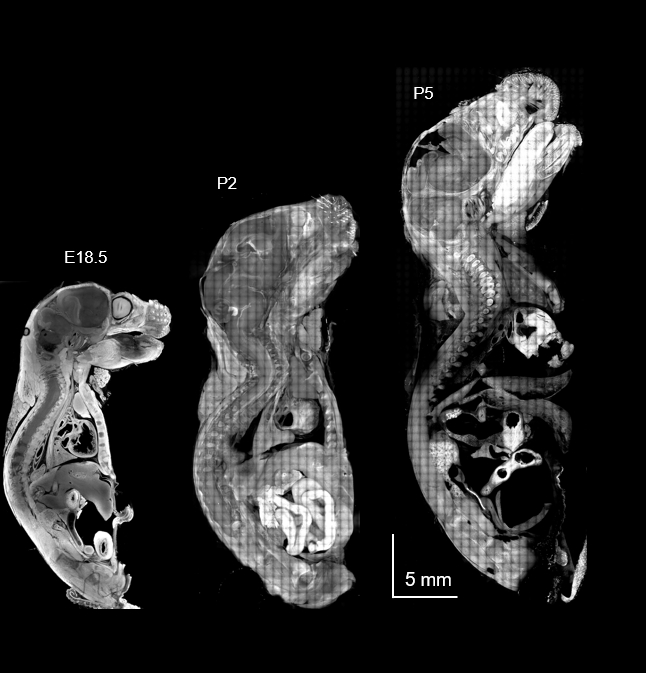
We develop chemical and materials strategies that expand the capability of molecular imaging. This includes expansion microscopy, anchoring and labeling chemistry, hydrogel engineering for ExM, and molecular processing methods that improve structural fidelity, signal accessibility, and multiplexing performance. Our focus is not on hydrogels as a standalone topic, but on chemistry and materials engineering that make advanced imaging more robust, versatile, and quantitative.
By engineering the chemical environment of biological samples, we aim to improve how biological structures are preserved, expanded, labeled, and measured. These efforts help bridge nanoscale structural imaging with scalable workflows that can be applied across diverse tissues and experimental settings.
Molecular Imaging and Neurodevices
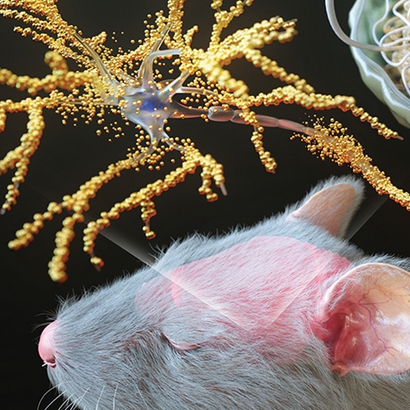
We investigate how biological structures can be transformed into functional devices. In particular, we engineer neuronal architectures through surface patterning, molecular perturbation, and controlled culture, then analyze their structural features using advanced imaging methods such as expansion microscopy and multiplexed imaging. These structural readouts are subsequently linked to metallization, electrical characterization, and device behavior.
Our long-term goal is to understand how biological architecture gives rise to function and to use that knowledge to create new neurodevice platforms. By connecting neuronal network design, molecular control, structural imaging, and electrical measurements, we aim to build bio-derived systems in which morphology is not merely preserved, but actively harnessed as a determinant of device performance and neuromorphic function.